Desktop app setup: quick start
This guide covers the two quickest ways to start using Platforma, right from the Desktop App. These methods are ideal for trying out Platforma, running small analyses, or when you don't need a permanent, shared backend service.
Method 1: use the built-in backend
This is the simplest way to get started. When you run the Desktop App for the first time, it gives you an option to run a temporary backend directly on your computer.
How to use it��
- Download Platforma Desktop for your operating system (macOS, Windows, or Linux).
- Launch the application.
- On the welcome screen, select the option to start a new project with the built-in backend.
- License Key: Paste your Platforma license key. You can get a license from platforma.bio/getlicense.
- Click Start.
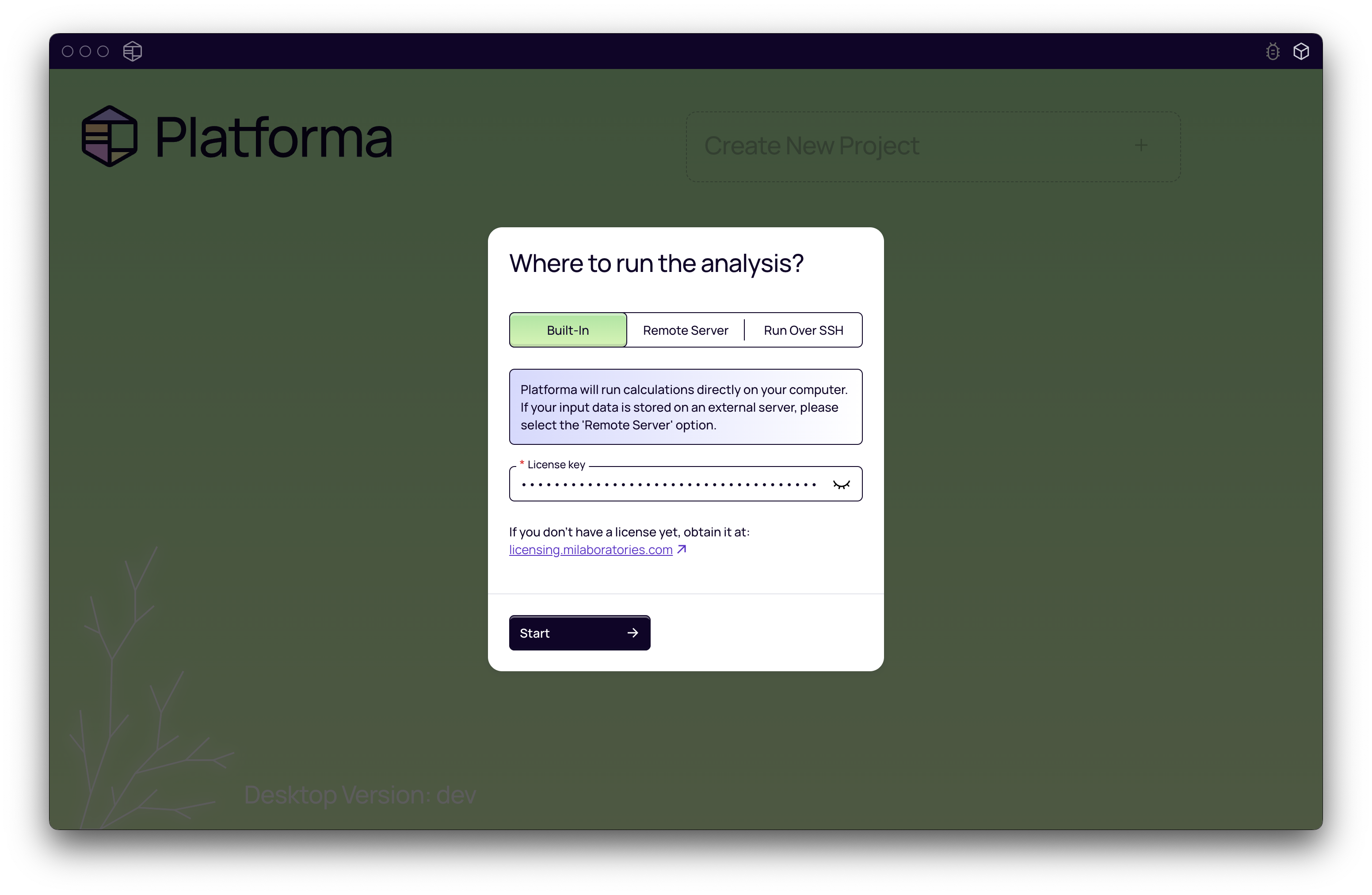
That's it! Platforma will start a local, temporary backend process for your session.
Limitations to be aware of
While incredibly convenient, the built-in backend has some important limitations:
- Resource consumption: all computations run on your local machine (your laptop or desktop). This will use your computer's CPU and RAM, which might slow down other applications.
- Not for large data: This setup is not designed for large-scale or long-running analyses. It's best suited for smaller datasets and exploring Platforma's features.
- Microsoft Windows compatibility: Many bioinformatics tools packaged as Platforma blocks are not compatible with Windows. A Linux-based server is the most compatible environment.
- It's temporary: The backend process stops when you close the Desktop App. No data is lost, but the backend is not running continuously.
Method 2: run over SSH
This option provides a great balance of convenience and power. It allows you to use the Desktop App on your local machine while running all the heavy computations on a more powerful remote server you have access to.
Platforma handles the technical details for you. It connects to your remote server, installs the backend, and starts it automatically.
When to use it
- You have SSH access to a more powerful Linux server (e.g., a university cluster or a lab server).
- You want to run analyses on larger datasets without bogging down your local machine.
- You don't want to manually install or manage a backend service.
How to use it
- On the Desktop App's welcome screen, click Run Over SSH.
- Host: Enter the server's address (e.g.,
myserver.example.comor an IP address192.168.1.100). If your SSH service runs on a non-standard port, you can specify it likemyserver.example.com:2222. - Login: Enter your username for the remote server.
- Authentication: The app will automatically detect which authentication methods are available. Select your preferred method (e.g., password or private key) and enter your credentials.
- License Key: Paste your Platforma license key. You can get a license from platforma.bio/getlicense.
- Click Connect.
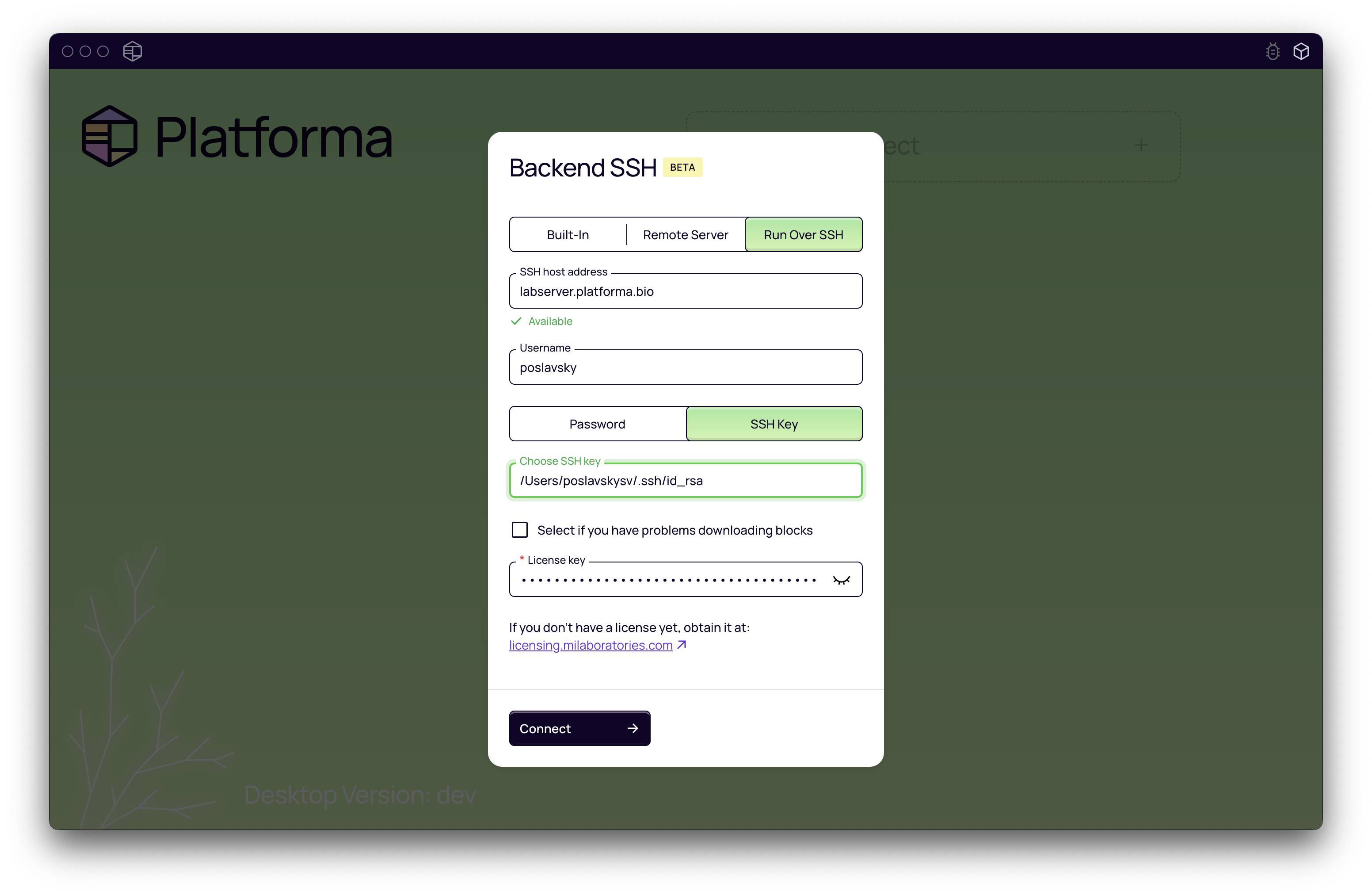
The Desktop App will now connect to the server and set up the backend. This might take a few moments. Once connected, you can use Platforma as you normally would, but all the work will happen on the remote server.
How it works and how to reset
The application installs the Platforma backend into a directory within your user's home folder on the remote machine. It does not require root access.
If you want to remove this automatically-installed backend and all its data, you can use the Reset option in the SSH connection menu.
Resetting the SSH connection will permanently delete the backend instance from the remote server, including all projects and results created with it.
Next steps: running your first analysis
Now that you have Platforma running, you're ready to analyze some data!
In Platforma, analyses are performed using Blocks — pre-packaged tools for common tasks like V(D)J annotation, clonotype analysis, and differential gene expression. These are the building blocks of your scientific discovery pipeline.