Functional Assay Data Integration
Bridging the gap between high-throughput next-generation sequencing (NGS) and functional validation is a critical step in antibody discovery. This guide demonstrates how to use the Immune Assay Data block in Platforma to import results from functional assays (e.g., ELISA, SPR, cell-based assays) and associate this data with your antibody clonotypes identified from NGS.
This process allows you to annotate your antibody sequences with crucial metadata, such as binding affinity, potency, or other performance metrics. By integrating this data, you can directly visualize, filter, and select the most promising antibody candidates within the Platforma ecosystem.
You will learn
- How to use the Immune Assay Data block to import functional data.
- How to configure matching parameters to link assay sequences with NGS clonotypes.
- How to use the imported functional data in downstream analysis tools like the Clonotype Browser and Clonotype Space.
Prerequisites
- A Platforma project with clonotyped antibody sequences (e.g., from running MiXCR Clonotyping).
- A functional assay data file in CSV, TSV, or FASTA format. The file must contain at least two columns: one with the amino acid sequences of the tested antibodies and at least one other column with the associated functional data (e.g., a 'Hit' label, binding affinity values, etc.).
- Note: If you upload a FASTA file, it will be automatically converted into a table with
HeaderandSequencecolumns.
- Note: If you upload a FASTA file, it will be automatically converted into a table with
Example example-assay-data.tsv file:
| Sequence | Hit |
|---|---|
| QVQLVESGGGL... | Ty1 |
| QVQLQESGGGL... | Ty1 |
Step-by-step guide
This guide will walk through the process of annotating a VHH dataset with "hit" information from a functional screen.
Add the Immune Assay Data block
Navigate to your project. In the left-hand menu, click Add Block and select the Immune Assay Data block from the menu. This block is a universal tool for associating any metadata with antibody or TCR sequences.
Configure the settings
Once the block is added, click on it to open the settings panel.
- Dataset: Select the dataset containing your NGS clonotypes. In this example, we'll use a dataset that has already been processed by MiXCR:
Tyl-pd / Redefined clonotypes. - Clonotype sequences to match: Choose the sequence field from your NGS data that will be used for matching against the assay data. Since our example uses full-length VHH sequences, we will select
VDJRegionInFrame aa.- Note: The choice of sequence is critical. The block can match amino acid sequences to nucleotide sequences (and vice-versa) by automatically translating them. For shorter sequences like a CDR3, you will need to adjust the matching parameters accordingly.
- Assay data to import: Click to upload your functional assay file. We will use the
example-assay-data.tsvfile mentioned in the prerequisites. The block supports comma-separated (.csv), tab-separated (.tsv), and FASTA (.fasta, .fa) files. - Assay data columns:
- Assay sequence column: Select the column in your file that contains the antibody amino acid sequences. In our example, this is the
Sequencecolumn. - Assay data columns to import: Select the metadata column(s) you wish to import and associate with your clonotypes. We will select the
Hitcolumn.
- Assay sequence column: Select the column in your file that contains the antibody amino acid sequences. In our example, this is the
Set the matching parameters
These parameters define how the platform will match the sequences from your assay file with the clonotypes in your NGS dataset.
-
Alignment score: Choose the algorithm for scoring the similarity between sequences.
- BLOSUM (
alignment-score): (Recommended) Uses a substitution matrix that accounts for the biochemical properties of amino acids. This is useful for identifying functionally similar, but not identical, sequences. - Exact match (
sequence-identity): Only considers sequences that are 100% identical.
- BLOSUM (
-
Score threshold: Set the minimum similarity score required for a match, ranging from 0.1 to 1.0. The default is
0.9, but for matching long, full VHH regions, we will increase it to0.95(or 95% similarity).tipThis value is highly dependent on the length of the sequence you are matching. For short sequences like CDR3s, a lower threshold may be necessary.
-
Coverage threshold: This defines the minimum percentage of the shorter sequence that must be covered by the alignment. We will leave this at the default of
0.95.
Run the analysis
After configuring all parameters, click the Run button. The block will execute, matching the sequences and annotating your clonotypes.
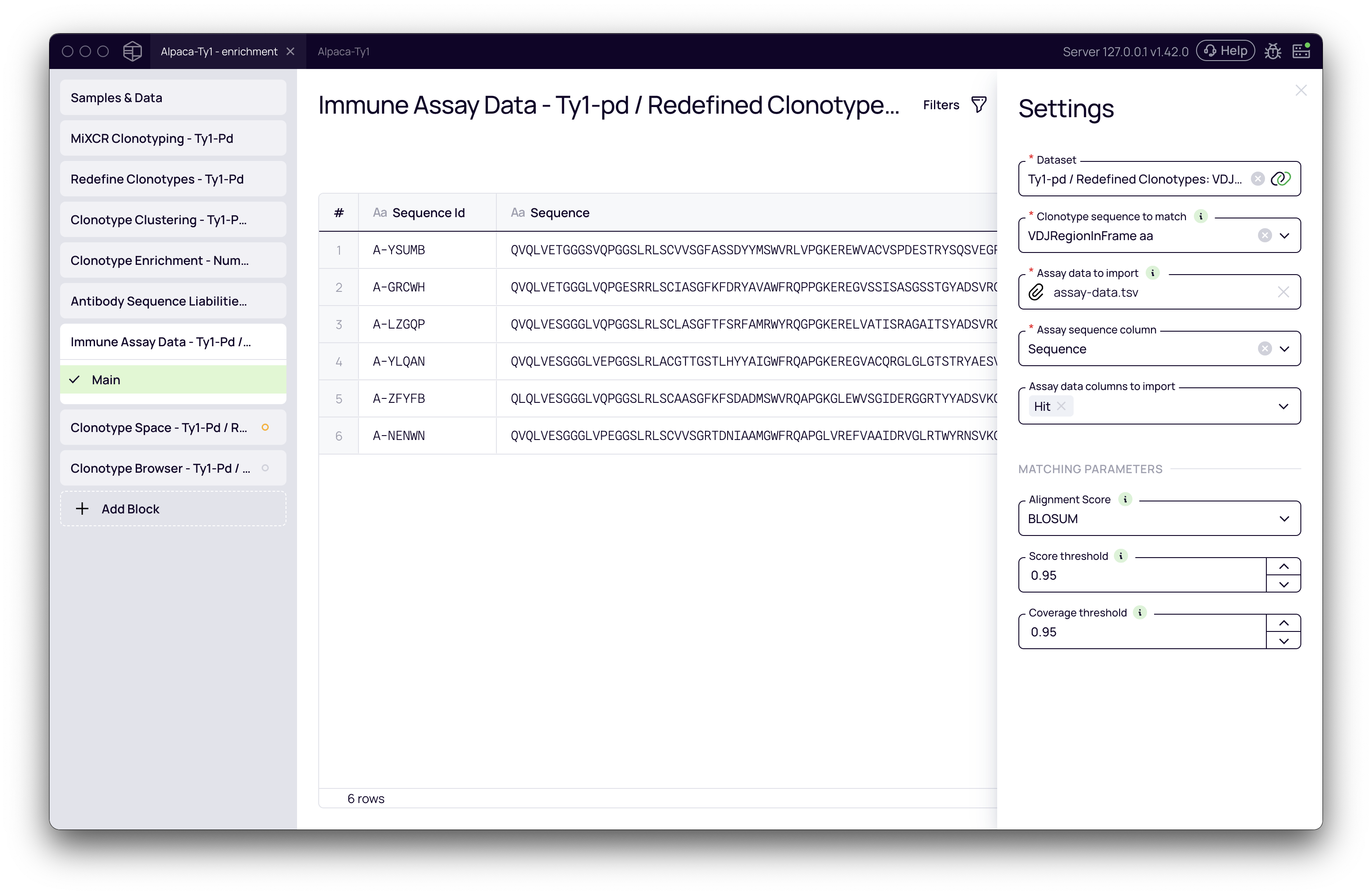
Understanding the results
Once the analysis is complete, the Immune Assay Data block displays a results table. This table lists each sequence from your input assay file and shows the number of Matched Clones found in your NGS dataset for that sequence.
More importantly, the functional data is now available as a new metadata column throughout your project for all matched clonotypes.
Downstream analysis
The integrated functional data can be used in various downstream applications to filter and visualize your top candidates.
Clonotype browser
Open the Clonotype Browser. You will now see your imported metadata column (e.g., Hit) available. You can filter and sort your clonotypes based on this functional data. For example, you can view only the clonotypes that were identified as hits in your assay.
Clonotype space
The Clonotype Space block allows for powerful visualization of your antibody repertoire. You can now use the imported functional data to color or highlight clonotypes on the UMAP plot.
- Open the Clonotype Space block.
- In the Data Mapping panel on the right, find the Highlight section.
- Select your new metadata field (
Hit). - Clonotypes that have been successfully annotated with this "hit" information will now be highlighted on the plot, allowing you to instantly see where your top functional candidates cluster within the broader antibody space.
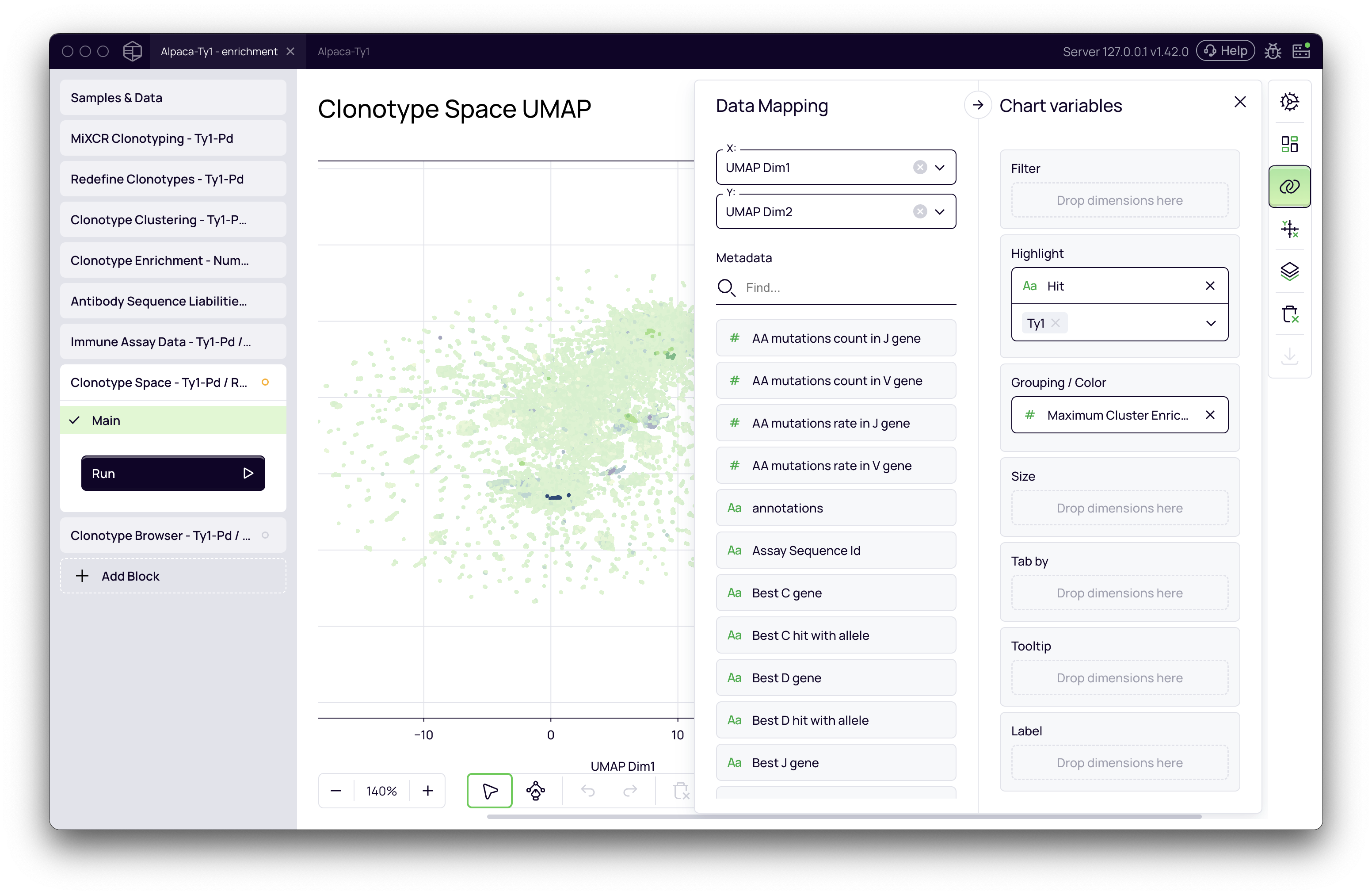
By combining the Immune Assay Data block with visualization tools, you can rapidly identify enriched clusters of functionally active antibodies, accelerating your lead selection process.