Types of Antibody Libraries
The first step in any antibody discovery project is understanding your source material and sequencing strategy. Whether you are mining natural immune responses from an immunized animal or screening a synthetic library, the structure of your antibody constructs and the way they are sequenced fundamentally shape the data you will analyze.
Platforma is designed to handle the full spectrum of antibody discovery data. This guide provides an overview of the most common library types and sequencing approaches, helping you identify your data format and understand how Platforma's specialized tools turn raw sequence reads into actionable biological insights.
Natural immune repertoires
Natural repertoires, sourced from B-cells of immunized animals (like mice, rabbits, or llamas) or humans, are the gold standard for discovering therapeutic antibodies. The process of in-vivo affinity maturation acts as a natural quality control, yielding candidates with superior stability and fewer developability issues. Next-generation sequencing (NGS) allows us to digitize and analyze these repertoires at immense depth.
Single-cell V(D)J sequencing
For traditional antibodies, which consist of a heavy and a light chain, preserving the native pairing of these chains is critical for retaining binding specificity. Single-cell technologies solve this by isolating individual B-cells and sequencing their paired heavy (VH) and light (VL) chain variable regions.
This approach, famously used in 10x Genomics V(D)J kits, provides a high-resolution view of the immune response, capturing not only sequence information but also cell-specific context. The data is tagged with cell barcodes (CB) to identify the cell of origin and unique molecular identifiers (UMI) to correct for amplification bias and ensure accurate quantification.
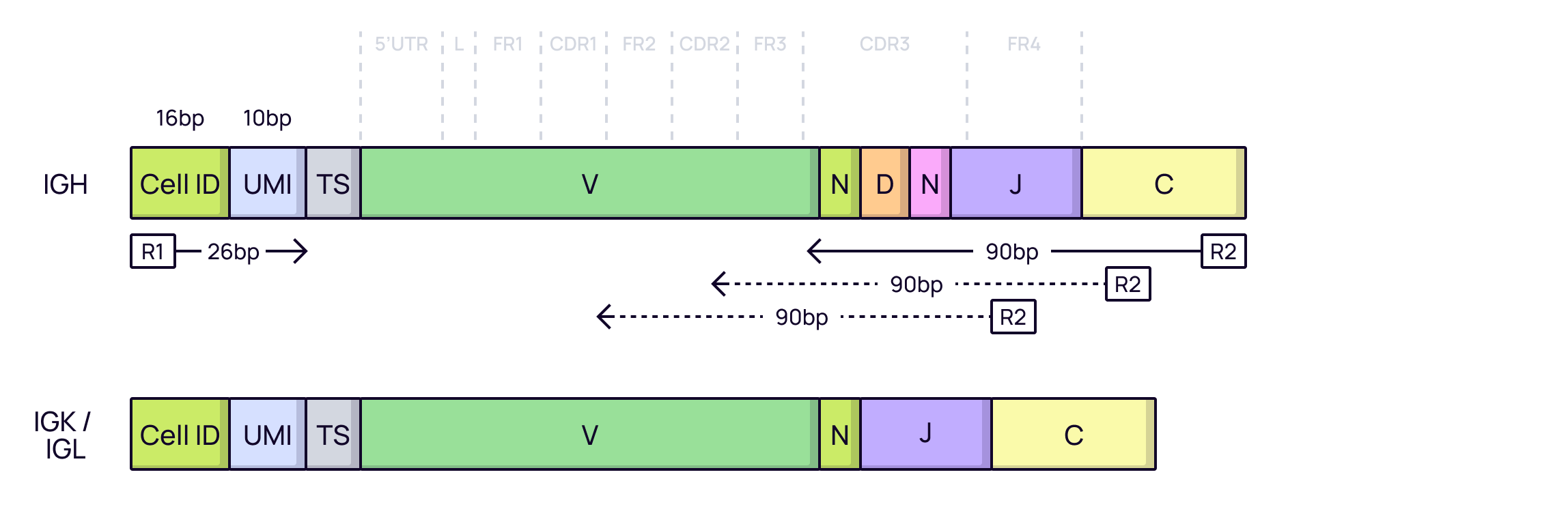
In Platforma: Data from single-cell platforms is the ideal input for the
MiXCR Clonotypingblock, which accurately assembles full-length V(D)J sequences and provides natively paired chain information.
VHH (Nanobody) sequencing
VHHs (or nanobodies) are single-domain antibodies derived from camelids (llamas, alpacas). Their small size, high stability, and ability to bind unique epitopes make them a revolutionary format for drug design. Since they consist of only a single heavy chain, the pairing issue is non-existent, and they are often analyzed from bulk RNA.
Sequencing can be done with or without UMIs.
-
Without UMIs: A straightforward approach where VHH sequences are amplified and sequenced directly. Quantification is based on raw read counts.
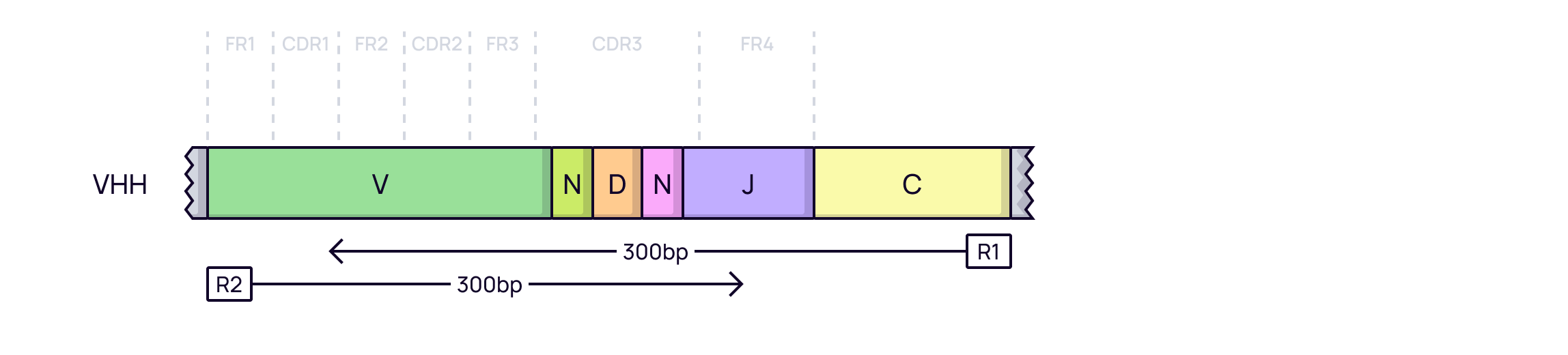
-
With UMIs: UMIs are added to each RNA molecule before amplification. This allows for precise, amplification-bias-free quantification of each starting VHH molecule, leading to more reliable results.
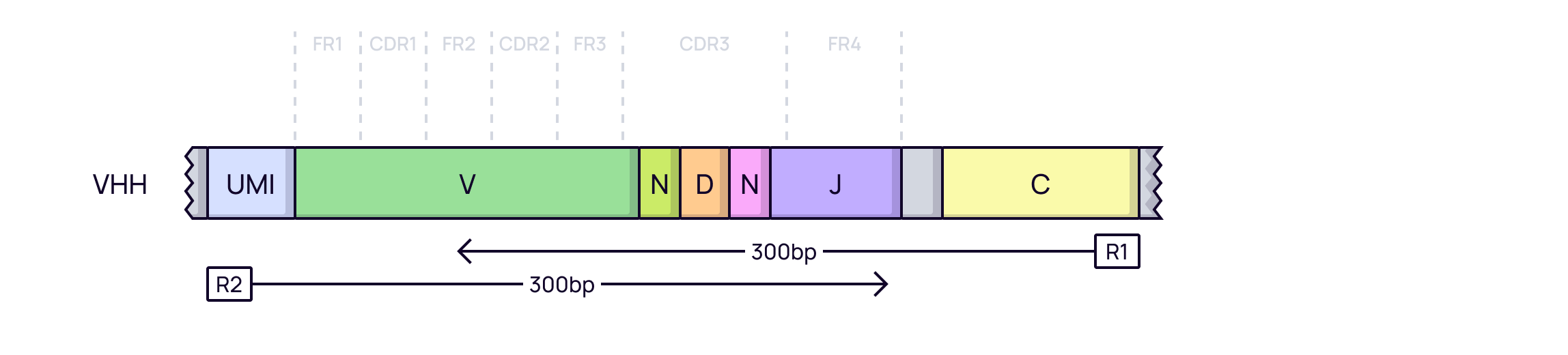
In Platforma: Both UMI and non-UMI VHH datasets can be processed using the
MiXCR Clonotypingblock. The workflow automatically handles UMI correction for highly accurate clonotype quantification.
Synthetic antibody libraries
Artificial (or synthetic) libraries, often screened using in-vitro methods like phage display, offer unmatched speed and versatility. They are essential for generating binders against challenging targets (e.g., toxic or non-immunogenic molecules) and for creating custom antibody formats.
scFv from display libraries
A common format for artificial libraries is the single-chain variable fragment (scFv), where the VH and VL domains are joined by a flexible linker. In a display campaign, NGS is used to quantify the enrichment of specific scFv clones after each round of selection (panning). The sequencing strategy depends on the read length.
-
Short-read sequencing (Illumina): This is the most common approach. Sequencing is often designed to cover the most diverse regions of the scFv, such as the heavy chain CDR3, which is sufficient to track clonal enrichment.
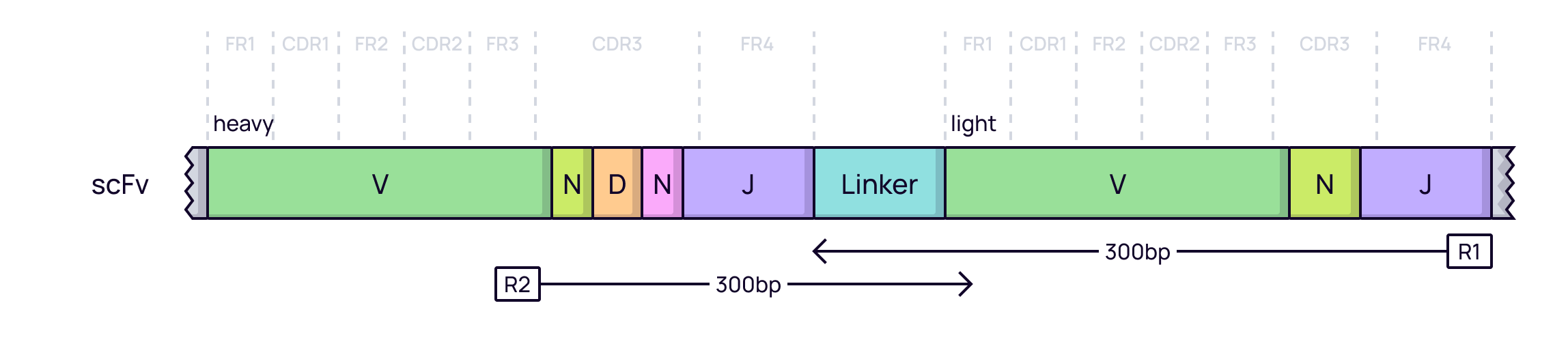
-
Long-read sequencing (PacBio / ONT): When the full sequence of the VH-linker-VL construct is required, long-read technologies can sequence the entire scFv cassette in a single pass.
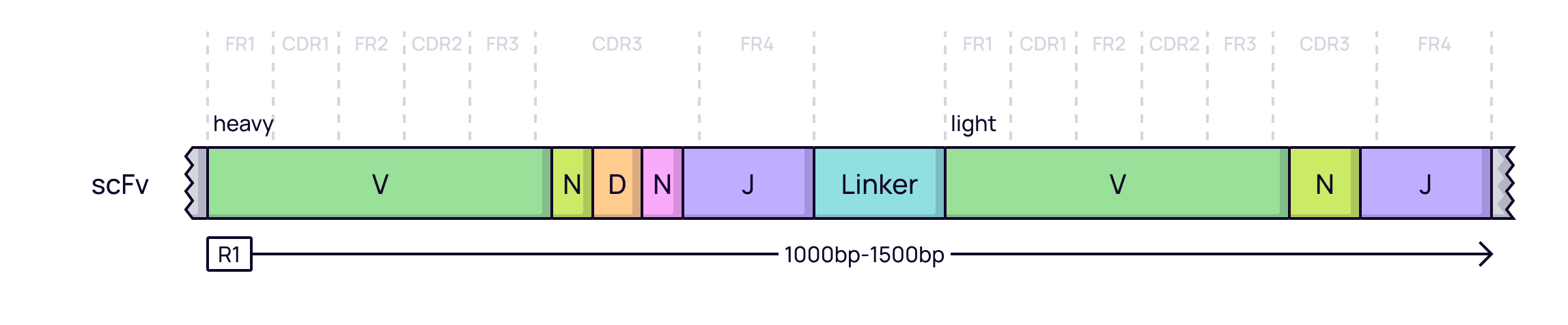
In Platforma: The platform provides specialized tools for different scFv workflows. The
MiXCR scFv Clonotypingblock is optimized for analyzing short-read data from scFv libraries.
Bulk synthetic libraries
In addition to scFv, discovery campaigns often use libraries built on standard antibody backbones (e.g., IgG) where diversity is introduced by randomizing specific regions like CDRs or even parts of the framework. These libraries are typically sequenced in bulk to assess diversity and identify clones with desired characteristics.
In Platforma: For this type of data, the
MiXCR Amplicon Alignmentblock is the appropriate tool. It is designed to handle bulk sequencing data from libraries with artificial diversity, allowing you to align reads to a reference backbone and analyze the randomized regions.
Summary
The choice of discovery platform and sequencing technology dictates the structure of your data. The table below summarizes the library types and points you to the right tools in Platforma for analysis.
| Library Type | Typical Source | Key Data Features | Sequencing | Recommended Platforma Block |
|---|---|---|---|---|
| Paired Heavy & Light Chains | In-vivo (mouse, human) | Natively paired VH/VL, Cell Barcodes, UMIs | Short Read (Illumina) | MiXCR Clonotyping |
| VHH (Nanobody) | In-vivo (camelid) | Single heavy chain domains, optional UMIs | Short Read (Illumina) | MiXCR Clonotyping |
| scFv | In-vitro (Phage Display) | VH-Linker-VL construct | Short Read or Long Read | MiXCR scFv Clonotyping |
| Bulk Artificial Library | In-vitro (Yeast/Phage Display) | Randomized CDRs/Frameworks on IgG backbone | Short Read (Illumina) | MiXCR Amplicon Alignment |
Next steps
Now that you have identified your library type, the next step is to process your raw sequencing data to identify and annotate the antibody sequences.
- Proceed to the Antibody Annotation guide to learn how to use Platforma's clonotyping tools.